Bacteriophages, often referred to as phages, offer promising and potent solutions for combating bacterial infections that have become resistant to antibiotics. The rise of antibiotic-resistant bacteria poses a substantial and pressing threat to global public health, affecting approximately 70% of bacteria responsible for causing infections as they become resistant to one or more antibiotics.
The development of new antibiotics to combat this challenge is both a costly and time-consuming endeavor, and it remains susceptible to the rapid development of bacterial resistance. In contrast, while bacterial resistance to phages can emerge, it does not seem to propagate through lateral gene transfer. Additionally, phages themselves can adapt through mutation to regain their ability to infect bacteria.
What makes phages particularly advantageous is that they have been identified for all known bacteria, allowing for the selective targeting and elimination of pathogenic bacteria strains.
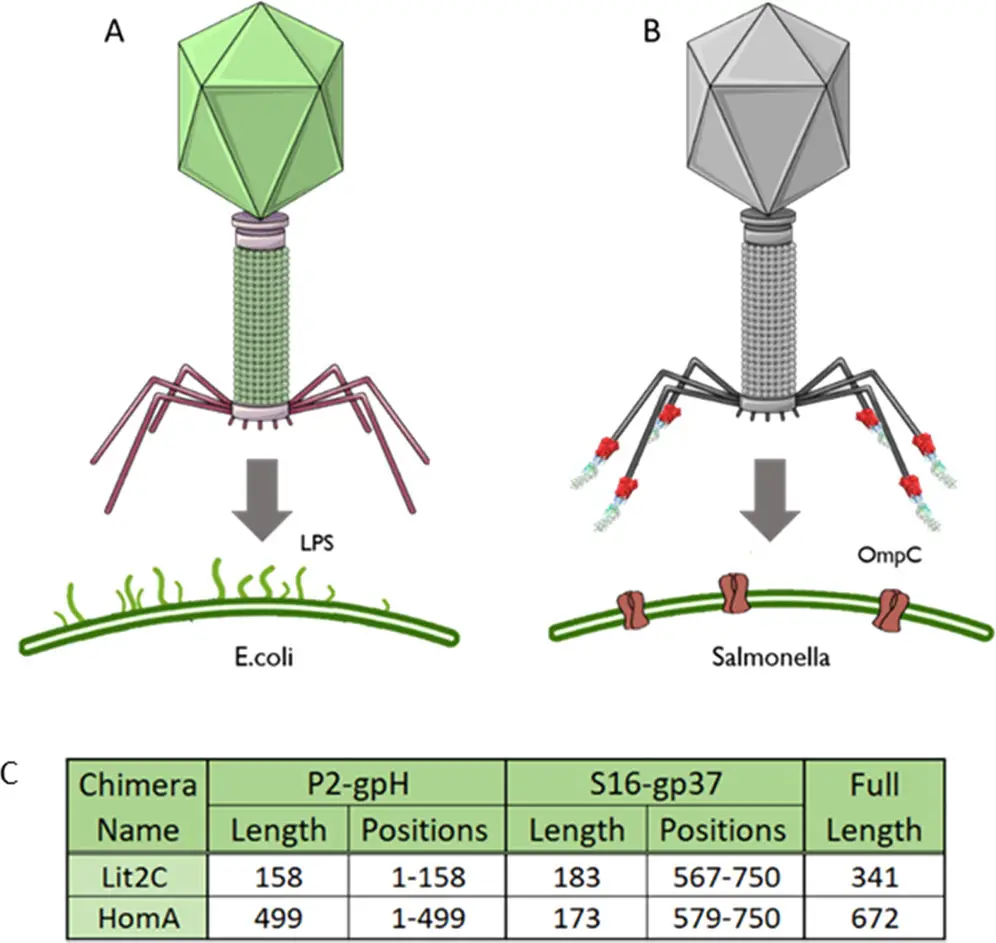
Scientists from Cardiff University and the University of Warwick undertook a remarkable project to modify the Escherichia coli phage P2, redirecting its specificity towards the pathogenic bacteria Salmonella. Through the application of genetic engineering, they reconstructed the tail fiber of the bacteriophage, a process known as bacteriophage pseudotyping. This term, bacteriophage pseudotyping, may be unfamiliar to many but entails the incorporation of a “foreign” viral envelope glycoprotein to change the bacteriophage’s targeting capabilities. In this case, it enabled the phage to infect a different type of bacteria than its original target, thereby expanding its host range.
The researchers explored two chimeric tail fiber designs, incorporating tail fiber genes from P2 and another phage, S16, which is known to infect and lyse Salmonella bacteria. Their objective was to refocus P2 towards a specific protein receptor present in Salmonella bacteria, a recognized pathogen. The selection of this target was motivated by the existence of a fully functional truncated S16 tail fiber.
Pseudotyping is not limited exclusively to bacteriophages; it can apply to various types of viruses. This process involves the alteration of a virus by incorporating foreign envelope glycoproteins, which can influence its targeting abilities. This concept of pseudotyping can extend beyond bacteriophages, allowing for the transformation of other viruses as well.
Moreover, the term “transformation” in virology typically refers to a different process – the introduction of new genetic material into cells by exposing them to foreign DNA. This technique is used for various purposes, such as genetic engineering and the creation of genetically modified organisms. The process of pseudotyping, while it involves genetic modifications, is more focused on modifying a virus’s envelope proteins to change its tropism or targeting specificity.
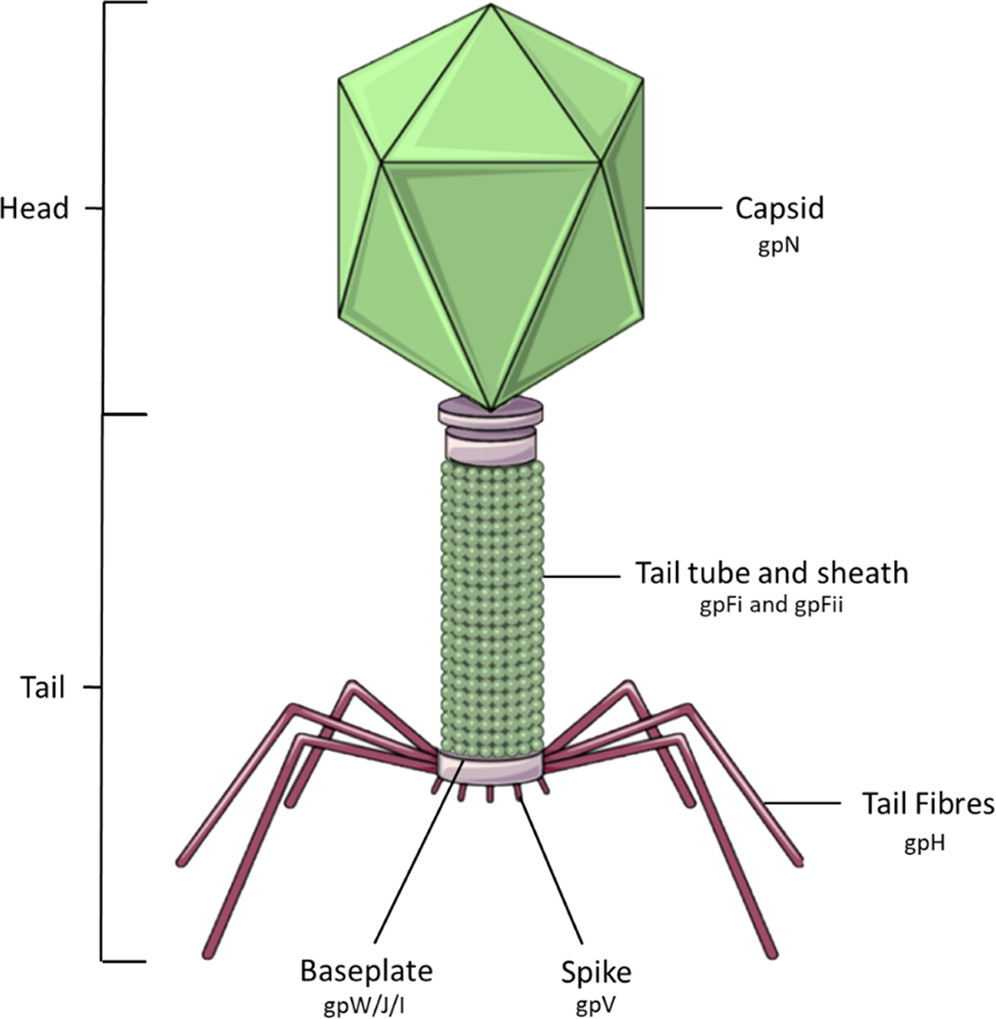
The phage
How did they do it?
The scientists devised two distinct chimeric tail fiber designs by utilizing different fusion points within the tail fiber genes from the two phages, P2 and S16. One approach was based on specific gene truncation points from these phages, while the second approach relied on a homology region identified through an alignment of the amino acid sequences of the tail fiber genes. This homology region spanned 17 amino acids and exhibited a 76% sequence identity.
To bring these designs to life, the researchers synthesized them as double-stranded DNA (dsDNA) and then amplified them using the polymerase chain reaction (PCR). The PCR products were subsequently subjected to treatment with a specific restriction endonuclease enzyme, which cleaved the amplicons at a specific site. Following this, the resultant plasmids were cloned and inserted into subcloning efficiency competent cells, a process facilitated by Invitrogen. To ensure the accuracy of the cloned sequences, the researchers confirmed their designs by performing Sanger sequencing on the plasmids.
Results
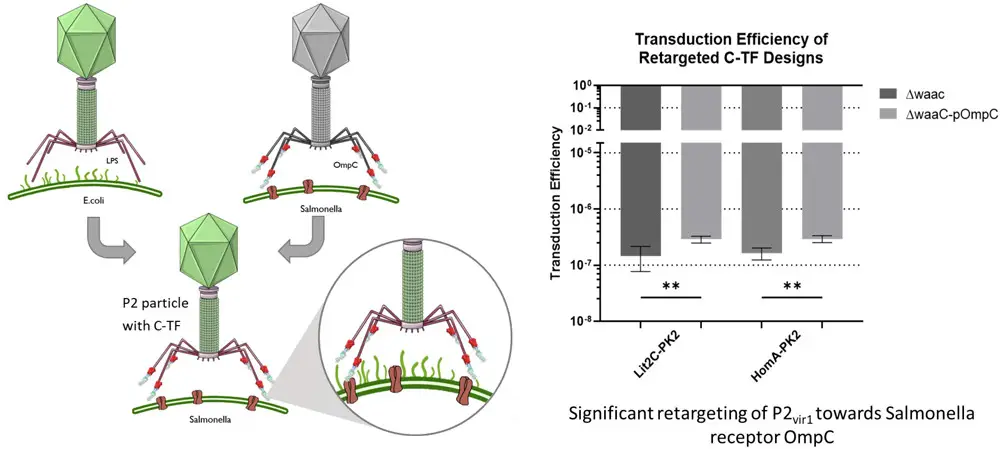
Chimeric tail fibers, formed by combining the P2 and S16 genes, were designed and created using two different approaches: one based on homology and the other on existing literature. By introducing these chimeric P2:S16 fibers onto the P2 phage, our findings indicate that the resulting phages were successfully retargeted away from their original P2 cellular target, the lipopolysaccharide, and were instead able to infect cells through the proteinaceous receptor, OmpC, which is the natural receptor for S16. This research provides compelling evidence that pseudotyping P2 is a viable strategy and can be employed to expand the range of hosts that P2 can target, particularly by using alternative receptors.
Engeneered phages were much better at infecting certain types of bacteria. They tested these altered phages on two different types of bacteria and found that they were about 2 times and 1.8 times better at infecting them. They also tested them on another type of bacteria that has a specific entry point (that was the phage manipulated target) for the phages, and they worked well on it too.
Furthermore, the possibilities stemming from this work extend to the potential creation of various chimeric tail fibers designed to target pathogenic bacterial threats. The engineering of P2 now allows adsorption through a foreign outer-membrane protein without the need for cultivation in its native host, offering a promising avenue for the development of designer phages to combat pathogenic bacteria, all guided by our knowledge of their surface proteome.
Reference and more credit
For more information about this article please find in the citation below Cunliffe, T. G., Parker, A. L., & Jaramillo, A. (2022). Pseudotyping bacteriophage P2 tail fibers to extend the host range for biomedical applications. ACS Synthetic Biology, 11(10), 3207-3215. NOTE all images published here are originally sourced from the same referenced study.
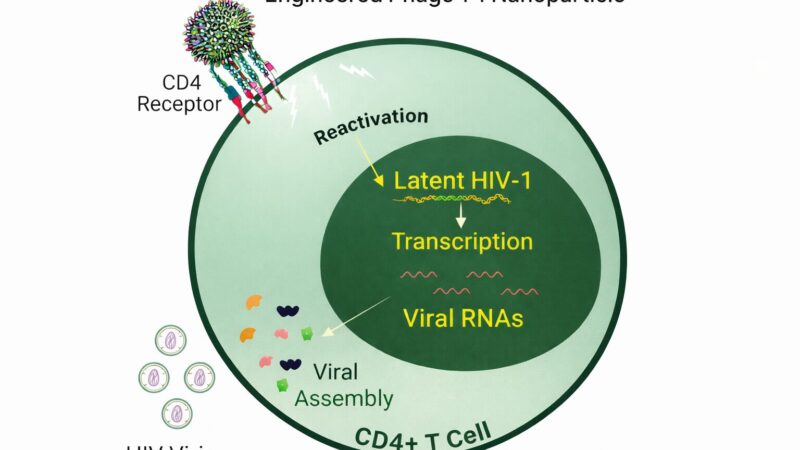

[…] of bacteriophage engineering, synthetic biology opens up a myriad of possibilities. Researchers can modify phage genomes to enhance their host range, enabling them to target a broader spectrum of bacterial pathogens. By incorporating genetic […]