
Bacteriophages, or simply phages, are viruses that infect and replicate within bacteria, playing a crucial role in shaping microbial communities and influencing bacterial evolution. As the field of bacteriophage research advances, the choice between traditional phage microscopy and next-generation sequencing (NGS) emerges as a pivotal consideration. In this comprehensive exploration, we delve into the nuanced interplay between these two techniques, dissecting their strengths, limitations, and the synergy they bring to the forefront of phage research.
The Power of Phage Microscopy:
Phage microscopy, with its roots tracing back to the dawn of microbiology, has long been the bedrock of understanding phage morphology, behavior, and interactions at the microscopic level. Through high-resolution imaging techniques such as transmission electron microscopy, researchers gain unprecedented insights into the structural intricacies of phages, witnessing the dynamics of infection and the delicate interplay between phages and their bacterial hosts. This level of detail is indispensable in the study of phage-host interactions, a cornerstone in phage therapy development and understanding microbial ecology.
Phage microscopy has traditionally played a pivotal role in the classification of bacteriophages. However, a significant shift occurred recently with the International Committee on Taxonomy of Viruses (ICTV) changing the approach to classifying these viruses. In the past, phages were categorized based on their shapes, often identified by informal names. The recent paradigm shift involves the assignment of binomial names, aligning with conventional scientific nomenclature. This updated classification process also takes into account genetic relationships rather than relying solely on morphological characteristics.
However, the microscopy has its limitations. While it excels in providing visual data, phage microscopy falls short in unravelling the genetic intricacies and broader genomic landscape of phages, leaving a crucial gap in our understanding of these dynamic entities.
Unlocking the Genetic Tapestry with NGS:
Enter next-generation sequencing, a revolutionary technology that has transformed the landscape of genomics. NGS enables the rapid and high-throughput sequencing of nucleic acids, offering an unprecedented ability to decode the genetic blueprint of organisms, including bacteriophages. This genomic approach has opened new vistas in phage research, allowing scientists to explore phage diversity, evolution, and functional genomics on a scale previously unimaginable.
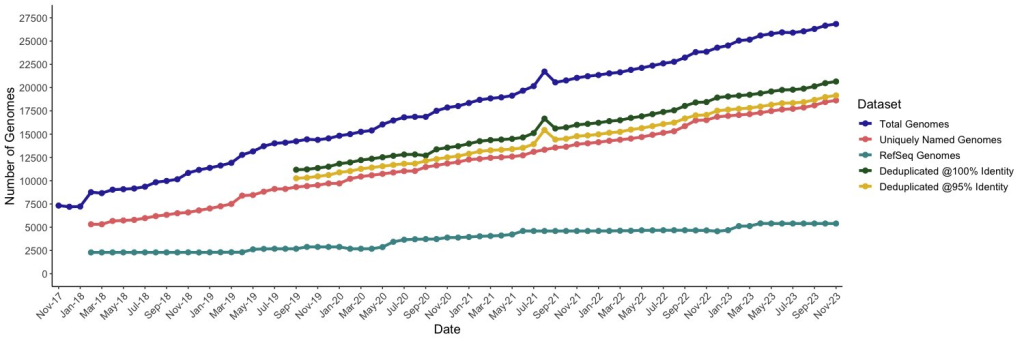
NGS not only accelerates the identification of phage genes but also sheds light on the broader genomic context, uncovering intricate relationships between different phage strains and their hosts. The wealth of genetic information obtained through NGS serves as a valuable resource for understanding phage biology, predicting their potential therapeutic applications, and deciphering the arms race between phages and bacteria.
Synergy in Simplicity: The Combined Force:
While phage microscopy and NGS might seem divergent, their combination forms a powerful synergy in bacteriophage research. Microscopy provides the qualitative observations and detailed morphology that NGS alone cannot capture. These visual cues guide researchers in selecting phages for genomic analysis, ensuring that the chosen strains are not only genetically interesting but also functionally relevant.
Recently, the synergistic capabilities of sequencing and microscopy have led scientists to make a remarkable discovery that would have been elusive through sequencing alone. Initially, perplexed by anomalies in their sequenced bacteriophage data, researchers suspected contamination. However, through microscopic examination, they unveiled an unprecedented finding: a satellite virus attached to another phage, behaving akin to a parasite. This groundbreaking observation, the first of its kind reported, was made possible by the collaborative integration of sequencing and microscopy technologies.
Conversely, NGS fills the gaps left by microscopy, offering a comprehensive genetic narrative that extends beyond individual phages to entire populations. This holistic approach is invaluable for understanding the diversity within phage communities, identifying conserved genomic elements, and discerning patterns that may govern phage-host interactions.
In the dynamic landscape of bacteriophage research, the choice between phage microscopy and NGS is not a matter of substitution but of integration. The judicious use of both techniques provides researchers with a multi-dimensional understanding of phages, combining the strengths of visual insights and genomic revelations. As technology continues to advance, the marriage of phage microscopy and NGS will likely yield even more profound insights, propelling the field toward new frontiers in microbial ecology, biotechnology, and phage therapy.


[…] The phage blog Can next-generation sequencing replace phage microscopy? […]
[…] their research without the added burden of operating and maintaining expensive equipment. Sometimes researchers may need to complement transmission electron microscopes with other services such as nex… to get more comprehensive answers for their […]