Recent advances in sequencing technologies have significantly improved the ability to generate large volumes of high-quality sequence data. However, the challenge lies in accurately assembling complete genomes from this data, a process that can be complex, particularly for phages, which often have small and highly variable genomes. Two primary methods are commonly used for genome assembly: de novo assembly and reference-based assembly. Each method comes with its own strengths and limitations, and the choice between them depends on various factors, such as the quantity and quality of the sequencing data, the availability of suitable reference genomes, and the specific goals of the research.
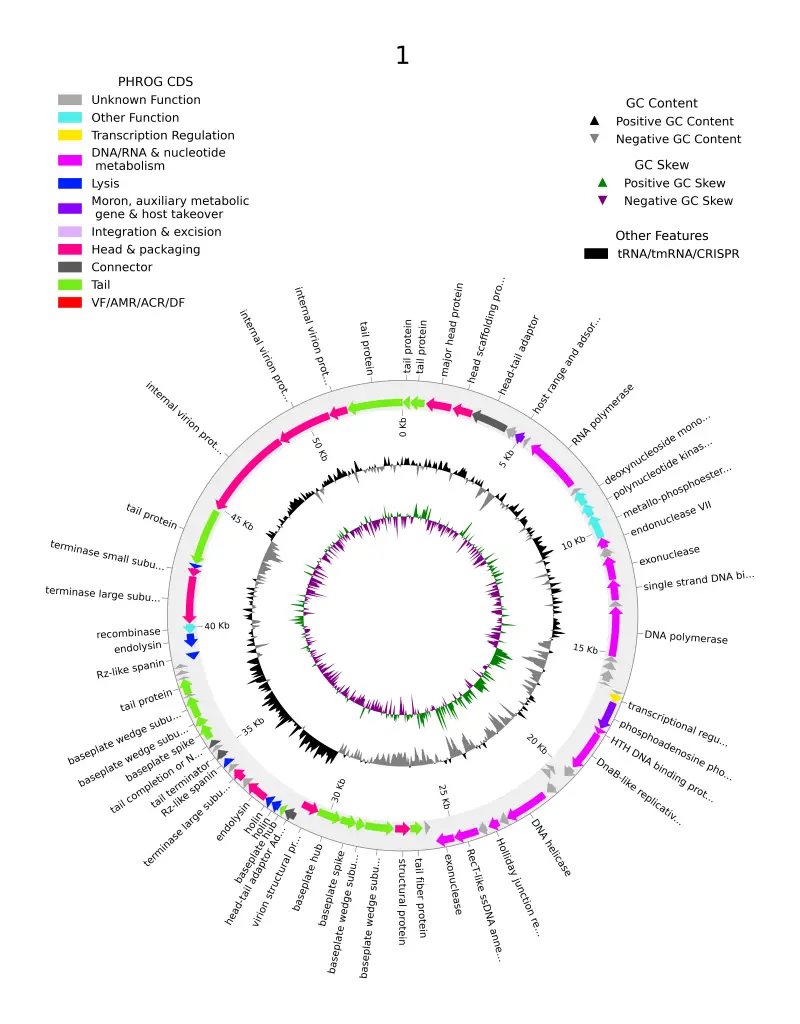
De-Novo genome assembly
De novo assembly is a method of genome assembly that involves piecing together overlapping fragments of DNA without the use of a reference genome. This method can be particularly useful for assembling phage genomes because phages are known to have highly variable genomes that may not have close relatives or well-annotated reference genomes. In such cases, de novo assembly can provide a more accurate representation of the phage genome, which can be used for downstream analysis and functional characterization.
De novo assembly can be a computationally intensive process that requires a high depth of coverage and quality sequencing data. It involves several steps, including quality control, error correction, assembly graph construction, and contig scaffolding. While the process can be time-consuming, recent advancements in sequencing technologies and computational algorithms have made it more efficient and accessible for many researchers.
Despite its strengths, de novo assembly also has some limitations. One of the main challenges is the risk of generating a fragmented or incomplete genome assembly, which can impact the accuracy of downstream analysis. Additionally, the method can be more prone to errors and misassemblies, which can be difficult to detect and correct.
This is an essential method for constructing phage genomes, particularly when a suitable reference genome is unavailable, which is often the case in phage genome assembly. The lack of reference genomes in databases is primarily due to the high diversity and small genome size of phages. This small size means that errors or biases introduced by reference-based methods can have a significant impact. Additionally, phage research is still in its early stages, and many phages remain unknown to scientists, further limiting the availability of reference genomes.
Reference-based genome assembly
Reference-based assembly is a genome assembly method that involves mapping sequencing reads to a previously assembled genome. This approach can be particularly useful for phage genome assembly when a high-quality reference genome is available. Unlike de novo assembly, which reconstructs the genome from fragmented sequences, reference-based assembly aligns sequencing data to an existing genome, making it a more straightforward and efficient process. Essentially, it provides the computer with a template, enabling the machine to generate a genome that closely resembles the reference.
Using a reference genome enhances the accuracy of the assembly and minimizes the risk of errors or misassemblies. Moreover, it offers insights into the genomic features of the phage, such as gene content and synteny, which are crucial for downstream analyses and functional characterization. Gene synteny, or the conserved arrangement of genes, is particularly relevant in bacteriophages, as their gene order is not random but organized in a way that can be understood. Therefore, with a suitable reference genome, it becomes easier to produce high-quality genome assemblies from sequencing reads.
However, the reference-based assembly also has some limitations. One of the main challenges is the lack of suitable reference genomes for many phages, which can limit the applicability of the method. Additionally, a reference-based assembly can be less accurate for phages with highly divergent genomes or regions of low coverage.
The use of reference-based assembly can prove advantageous for constructing phage genomes, but only when an appropriate reference genome is obtainable. However, given the significant differences that exist between phage genomes, finding a compatible reference genome is an infrequent occurrence. As such, it’s crucial that researchers diligently assess both the quality and completeness of the reference genome, as well as the appropriateness of the sequencing data, before beginning a reference-based assembly endeavour.
Optimal Approach for Assembling Phage Genomes
Assembling genomes for organisms such as bacteria and fungi can be effectively accomplished through a reference-based assembly approach. This is due in large part to the availability of numerous well-annotated reference genomes for bacteria that are publicly accessible. However, when it comes to phage genomes, highly variable sequences and a lack of close relatives or well-annotated reference genomes can make reference-based assembly challenging. As a result, de novo assembly is generally considered to be the preferred method for assembling phage genomes. Several software tools, including SPAdes and Shovill, are available for this purpose.
Another effective approach is hybrid assembly, which involves combining long and short sequencing reads to generate a high-quality genome. Tools like Tricycler can be used for this method. The advantage of using both long and short reads lies in their complementary strengths, which improve the overall assembly quality, though we won’t delve into the specific weaknesses of these technologies in this article.
It’s important to remember that there are numerous open-source and commercial genome assembly tools available, each offering distinct advantages and limitations. For a clearer understanding, you might want to check out a “Simplified Workflow to Assemble and Annotate Phage Genomes,” which provides a straightforward sketch and outlines the tools used in the process. If you have any questions, feel free to leave a comment or send me an email at raphael.hans@thephage.xyz.




Leave a Reply
You must be logged in to post a comment.