Overview
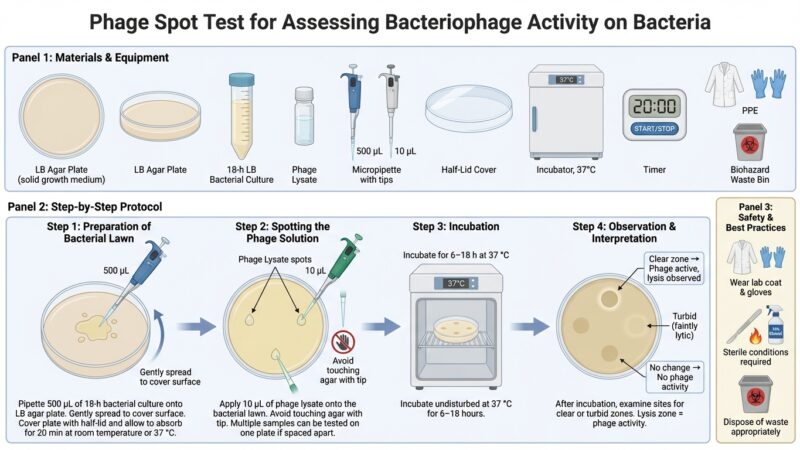
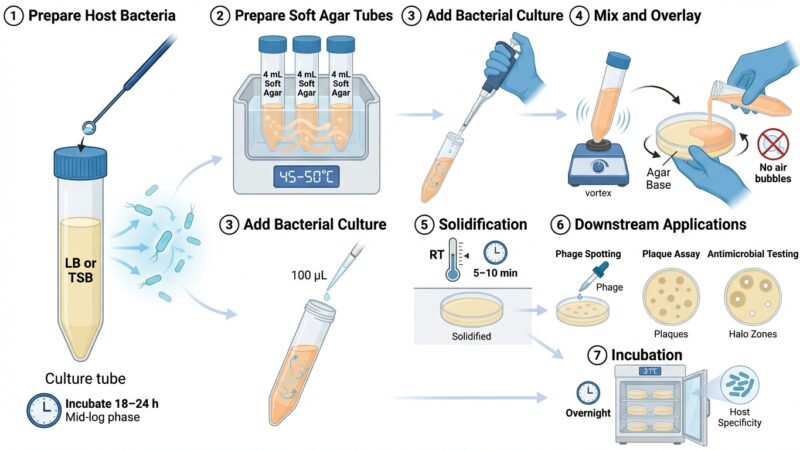
This protocol outlines the isolation and purification of bacteriophages from soil or sewage samples using bacterial host strains. Samples are enriched with host bacteria, followed by centrifugation and filtration to obtain phage-containing supernatants. Phages are detected using plaque assays with soft agar overlays.
Plaques (zones of lysis) indicate phage activity and are further purified through repeated plaque picking and re-plating. The final result is a filtered phage stock suitable for downstream applications such as titer determination and host range analysis.
Biosafety and Handling Notes
- Biosafety Level: Use BSL-1 or BSL-2 practices depending on host strains. Treat environmental samples as potentially hazardous.
- PPE: Lab coat, gloves, and eye protection required.
- Chloroform: Toxic; handle in a fume hood and dispose of as chemical waste.
- Waste Disposal: Autoclave biological waste; disinfect liquid waste as required.
Acronyms
SM – suspension media
LLB – Lennox LB
LLBA – Lennox LB Agar
TALLB- Top agar Lennox LB
MCF – Microcentrifuge
It is optimal to work with five strains at a time and five samples at a time. This means you will be working with 25 Falcon tubes per experimental day.
Reagents
- Soil or sewage sample – 50 g or 50 ml
- 50 ml falcon tubes – either bioreactors or flat lid with tape
- 2x strength LLB (sewage) or 1x LLB (soil)
- LLB agar plates
- LLB agarose top agar
- Suspension media
- Five target strains
- Pipettes and tips
- Serological pipettes
- 0.22 um filters
- 10 ml syringes
- Microcentrifuge tubes
- Sterile pasture pipettes and bulbs
- Water bath set to 55 °C for top agarose
- Chloroform
- 3 g Agarose
Methods
Day Zero
- Prepare overnight cultures of target strains, up to 5 per sample, in 10 ml LLB with a single colony. Incubate at 37°C with shaking at 225 rpm for 16 h.
- Prepare all of your tubes for D1 and all of your plates for D2
Day One (D1) Enrichment
- Distribute sewage (10 ml) or soil (10 g) samples between 50 ml falcon tubes. One tube per strain.
- Add 5 ml 2 x LLB to sewage sample or 10 ml LLB to soil sample.
- Add 1 ml SM to each tube
- Inoculate each tube with 100 µl of the desired target strain from the overnight culture.
- Gently vortex the tubes to mix all ingredients.
- Close or tape the tube lids down, incubate at 37 °C with 225 rpm shaking for 18 hr.
- Make new set of overnights for phage screen tomorrow with the same strains.
Day Two (D2): Filtration and Plating
- Remove overnight cultures and samples from the shaking incubator.
- Centrifuge sample Falcon tubes at 5,000 × g, 4 °C, for 45 min.
- Remove supernatant and filter-sterilize into labeled 15 mL tubes using a 10 mL syringe and 0.22 µm filters.
- For each sample, prepare two culture tubes. Combine:
- 100 µL overnight strain
- 100 µL filtered supernatant
- 25 µL SM
Store stocks at 4 °C.
- For each strain, prepare a negative control using SM only.
- Incubate culture tubes at room temperature for 20 min.
- Remove TALLB from the water bath; ensure it is molten and homogeneous. Add 3 mL TALLB to each culture tube using a serological pipette (pipette down the side to avoid bubbles).
- Pour onto LLBA plates and gently move in a figure-eight motion to spread evenly.
- Leave plates undisturbed until agarose solidifies (~30 min or less).
- Incubate plates upside down at 37 °C overnight.
- Prepare another set of overnight cultures with the same strains.
Plate calculations:
- 1 strain, 5 samples → (2 × 5 × 1) + (1 × 1) = 11 plates
- 3 strains, 5 samples → (2 × 5 × 3) + (1 × 3) = 33 plates
- 5 strains, 5 samples → (2 × 5 × 5) + (1 × 5) = 55 plates
Day Three (D3) Plaque Screening
- Remove overnight cultures from the incubator.
- Check plates incubated at 37 °C. If plaques are present, remove those plates; otherwise, continue incubation up to 48 h total.
Two possible outcomes:
1. Confluent lysis:
- Scrape agarose into a 50 mL Falcon tube and add 15 mL SM.
- Vortex to break up agarose (note: this is not a pure phage stock).
- Centrifuge at 5,000 × g, 4 °C, for 45 min.
- Filter-sterilize the supernatant into a new tube and store at 4 °C.
- Streak onto LLBA plates for single plaques.
- Mix 100 µL overnight bacteria + 25 µL SM + 3 mL TALLB and pour onto the plate. Do not swirl.
- Incubate overnight at 37 °C.
- Pick a plaque the next day and repeat twice.
2. Isolated plaques:
- Prepare an MCF tube with 500 µL SM and 20 µL chloroform.
- Use a Pasteur pipette to pick a single plaque and transfer it into the tube.
- Collect up to three plaques per type (based on morphology).
- Gently vortex and allow chloroform to settle. Store at 4 °C.
- Streak and plate as above.
- Incubate and repeat plaque picking twice more.
Day Four (D4)Confirmation and Stock Preparation
- Check D3 plates and any remaining D2 plates.
- If plaques appear on D2 plates, repeat D3 procedures; otherwise, dispose of plates appropriately.
- For purified plaques showing confluent lysis, prepare an axenic working stock as in D3. Store at 4 °C after filtration.
- Determine phage titer using a dilution series.
- For high-titer stocks (≥10⁹), consider liquid propagation (note: not all phages propagate efficiently in liquid). Start with small volumes before scaling up.
For more phage-related lab protocols, click here