What is phage typing?
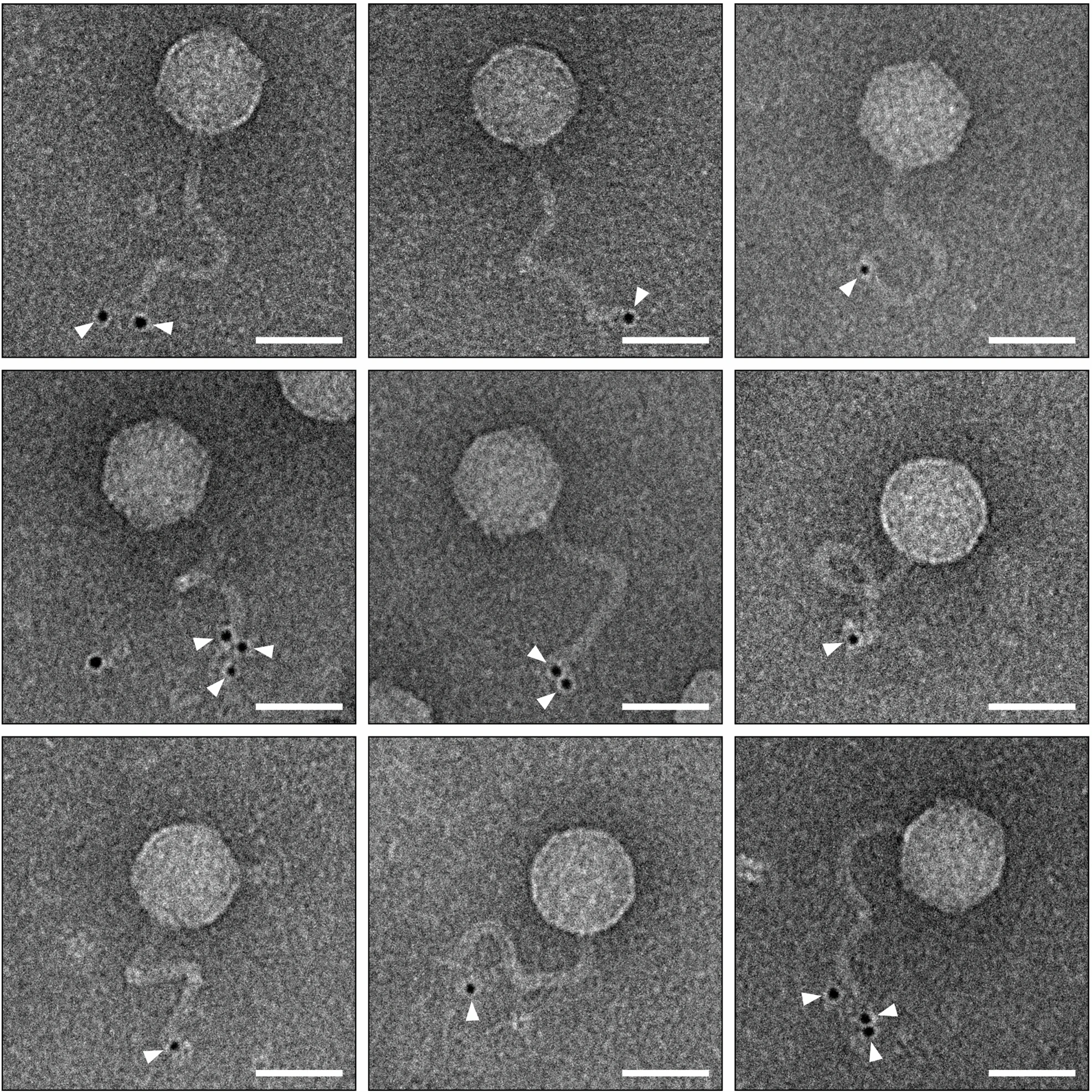
Phage typing is a quick and low-cost phenotypic method that uses bacteriophage specificity to detect and identify single strains of bacteria. Is simple to understand this concept once you know the phage life cycle. Bacteriophages use one of two mechanisms: lytic or lysogenic. In the first step, the phage inserts its genetic material into the cell, converting it into a factory for the production of more phages, which will eventually lyse the host. During lysogenic action, the phage’s injected genome is incorporated into the cell’s DNA, transforming it into a prophage that will generate progeny when reproduction conditions are favourable. During bacterial cell division, the phage nucleic acids multiply along with the bacteria’s genome.

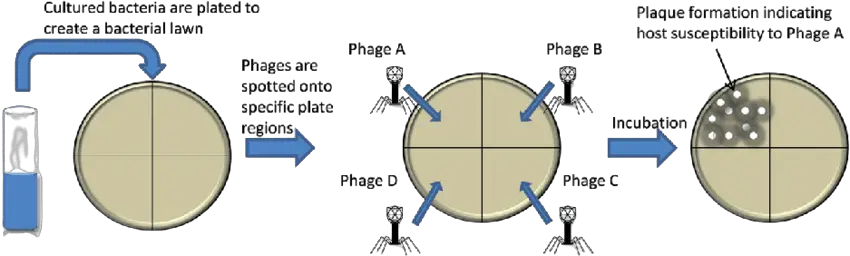
Furthermore, these bacterial viruses are highly specific, with some phages infecting only one strain of bacteria. Scientists take advantage of this strange ability by identifying the bacteria in question using lysis action on the bacterial lawn. By simply dividing a plate containing a lawn of investigated bacteria into a grid and seeding each zone with a panel of different types of lytic phages, the identity of the strain of bacteria can be determined by observing which portions of the panel show clear zones. The phage specificity discussed above can be used to classify bacteria based on phage type. Lytic bacteriophages are ideal for this purpose because the results are observable via cell lysis, which forms a plaque on the bacterial lawn. Bacterial strains within a specific serovar such as Salmonella enterica serovar Typhimurium can be classified into a number of phage types based on their susceptibility to lysis by a group of phages with varying specificities.
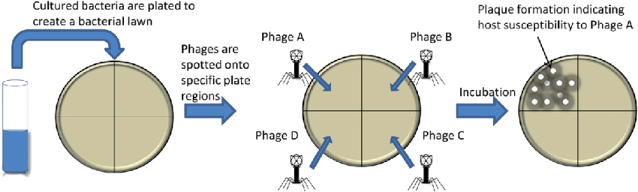
Phage typing procedure in detail
- Prepare Luria-Bertani (LB) agar plates by following the manufacturer’s instructions (I wrote a Detailed post on how to make an outstanding agar and LB agar plate preparation)
- Draw lines on the base side of the media plate to make separate portions where phages will be spotted. The number of divisions is determined by the number of phages used for typing; too many divisions may jeopardize the reading of results.
- Write the laboratory identity number or code of the bacteria to be typed on the agar plate. If you have more than one bacteria to type, make sure to use a separate agar plate for each.
- Prepare an agar overlay media. The overlay media is semisolid, made from broth mixed with 0.5-0.7% w/v agar.
- Measure 4 milliliters of your overlay in a sterile graduated falcon tube and add 100µl of overnight bacteria to be tested.
- Vortex the mixture briefly but properly
- Pour the mixture on the surface of the plate that corresponds with the label of the bacteria used. Tilt and swirl the plate slightly to ensure the overlay spreads evenly across the surface of the base media, then set it aside to solidify.
- Spot 10µl of the phage preparation on a plate with each phage having its own division with a corresponding label.
- Let the spot dry before shaking the plate, to avoid overlapping of plaques. Working under a biosafety cabinet will significantly reduce the waiting time.
- Incubate the plate for hours depending on the type of bacteria you are dealing with. Many of the common bacteria may require 37℃ for 24 hours.
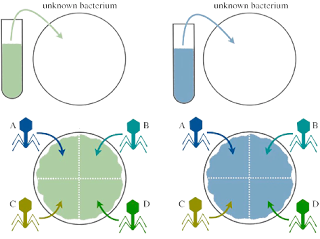
Reading results
After incubation examines the area where the phage was spotted for phage plaque. A clear plaque will be observed for all plate divisions where the bacteria were susceptible to phage, and no clearance will be observed where there was resistance. Before typing the bacteria based on their susceptibility, record the results as positive and negative (As seen in the figure below).
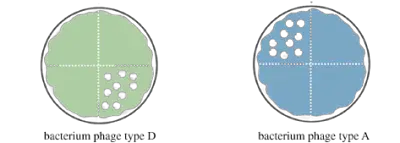
Important things to not when doing the procedure:
- The temperature for the overlay should not be too high to kill the bacteria, nor too low to allow the overlay to solidify.
- Plates should be left idle after being spotted with phages until they dry.
- The overlay should be spread smoothly because a rough surface will impede the results.
- Overlay that is about to solidify should not be used because it will not produce a smooth surface.
- Delays in swirling the overlay immediately after pouring it on the surface may result in uneven media distribution.
Applications of phage typing
In general, phage typing is used to identify and distinguish different strains of a given species that have been isolated from various sources (disease, food, water, environmental) or geographical locations or any other aspect that you can think of.
Specific application
- Phage typing is used by microbiologists to determine the interrelationship of bacterial species. Phage typing, in particular, offers a quick and low-cost method for epidemiological surveillance and determining the infection source in public health emergencies.
- Bacteriophage testing has long been considered the gold standard method for monitoring Salmonella Typhimurium and Salmonella Enteritidis. This system, for example, can distinguish over 300 definitives of S. Typhimurium based on their lysis patterns to a panel of Salmonella phages.
- It is frequently used for Escherichia coli, a type of bacteria that is part of the normal intestinal bacteria in both human and animal intestines and is particularly vulnerable to T-phage viruses, both ‘T-odd’ phages like T1 or T7 and, to a lesser extent, ‘T-even’ phages like.
- It is currently the most widely recognized typing technique for Staphylococcus and is still widely used for subdividing Pseudomonas aeruginosa serotypes.
- In some applications, such as contamination control in bioprocesses, phage typing remains a viable option available to the lab setting genotypic tools such as clustered regularly interspaced short palindromic repeats (CRISPR) typing, whole-genome sequencing, and so on.
Limitations of phage typing
Bacterial phage typing necessitates a multifaceted panel of prescreened and well-characterized phages, in addition to considerable technical expertise and strict control over the testing environment, as this can affect results. Only a few laboratories in the world have all of these features, as well as the means, facilities, and well-trained personnel to perform phage typing for any industrial application or bioprocess.
Although, given the world’s rapid technological advancement, this may become one of the most common tests performed in the near future.