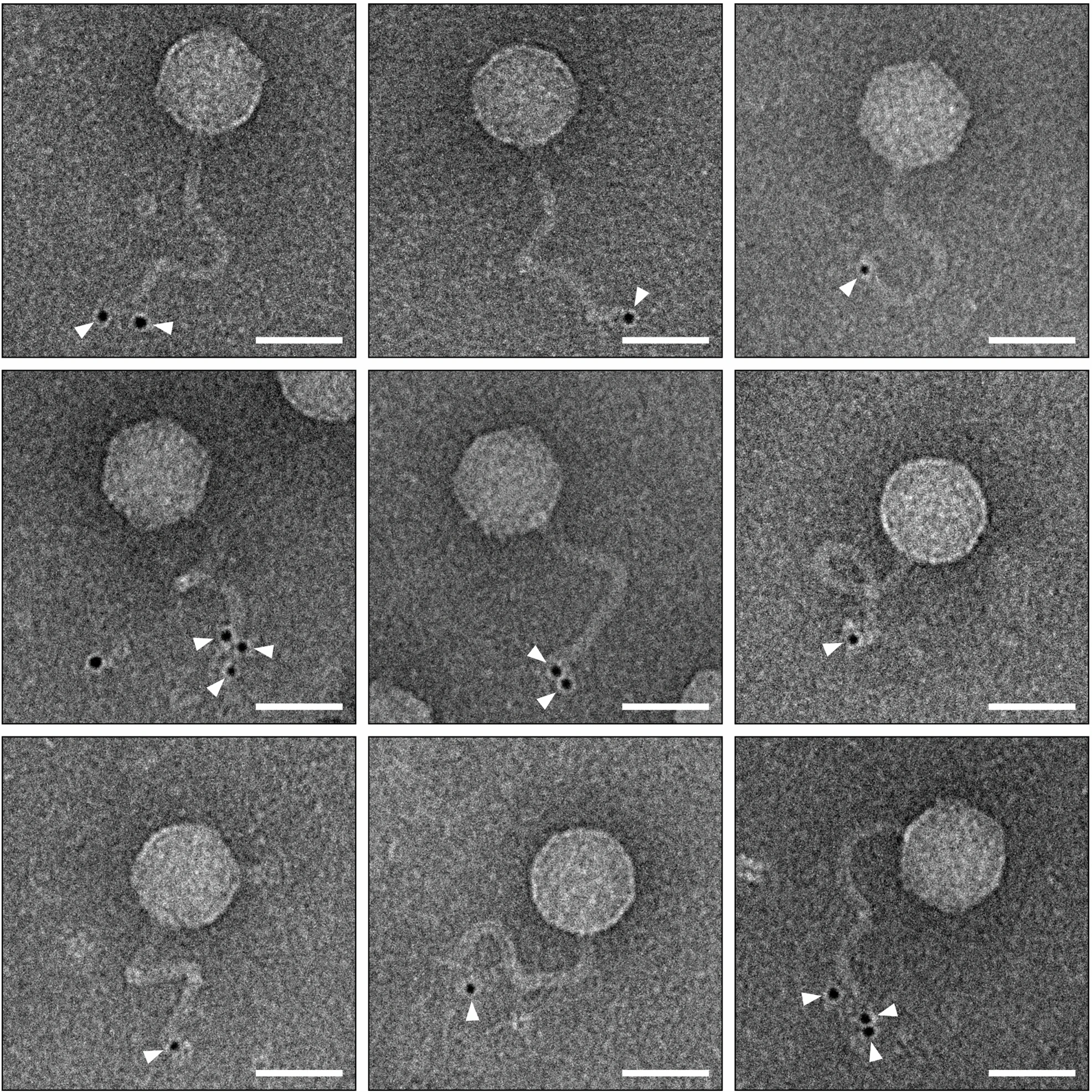
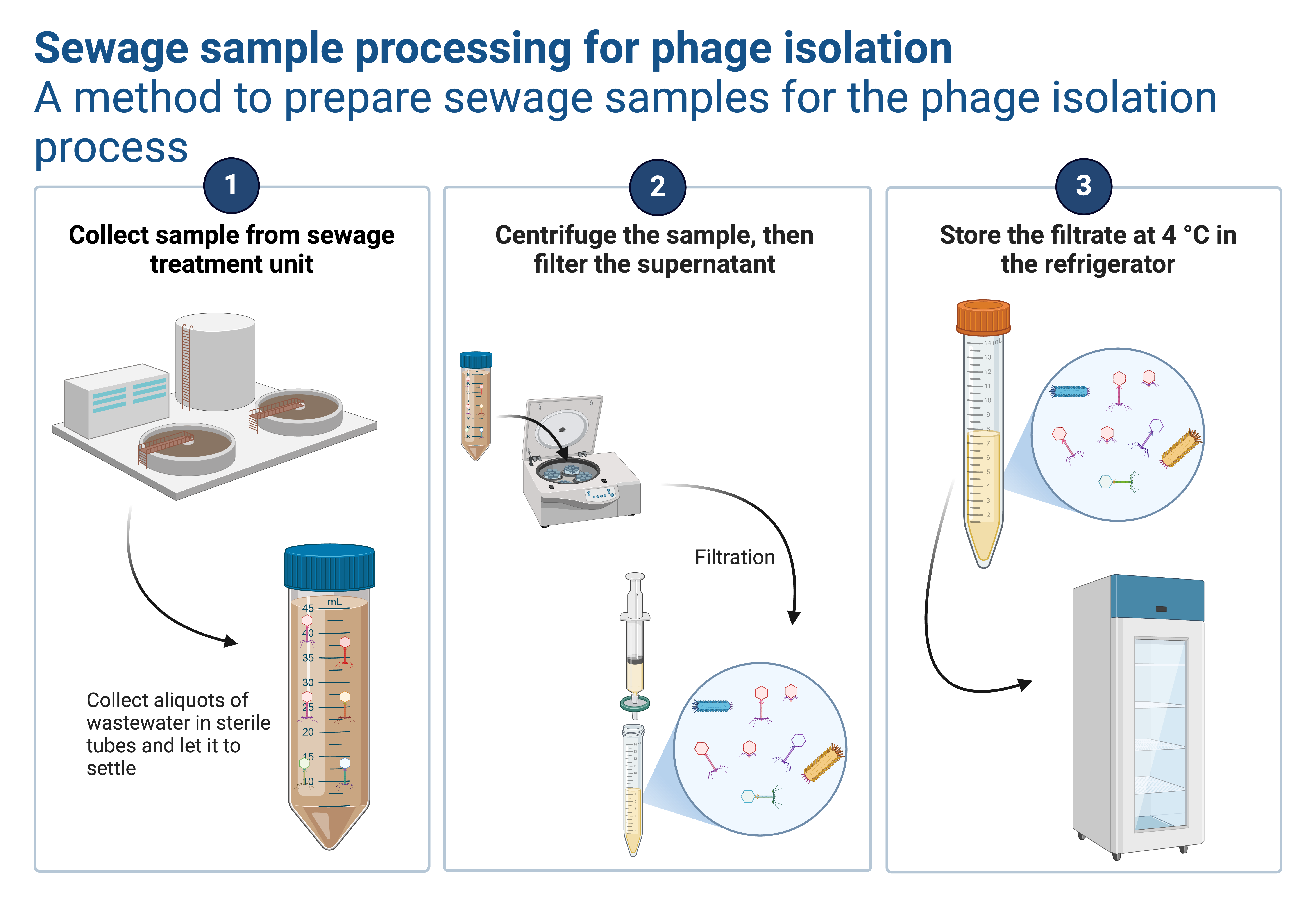
The one-step growth curve experiment determines the phage life cycle duration, including latent period, rise period, and burst size, by limiting infection to a single round. The procedure involves adsorbing phages to bacteria at a low MOI, removing unattached phages, diluting the mixture to prevent secondary infections, sampling over time, and plating for plaque enumeration. This protocol, adopted from Kropinski (2018) and Li et al. (2022) involve removing unadsorbed phages through centrifugation and dilution, bypassing the need for chemical agents like chloroform.
Materials
Biological materials:
- A high-titer phage preparation (≥10⁵ PFU/mL) (Protocol to quantify your phages can be found here. If you have done the labwork already, here is the PFU calculator to help you work out phage quantity)
- Host bacterial strain
Consumables:
- LB broth (or appropriate medium)
- LB agar plates (1.5% agar)
- Soft agar (0.6–0.7%) (Learn how to make soft agar here)
- SM buffer or PBS (Learn how to prepare SM buffer here and how to prepare PBS here)
Equipment:
The following are the major pieces of equipment needed (Small lab equipment has been omitted from the list)
- Incubator
- Centrifuge (≥10,000 × g)
- Spectrophotometer (for OD₆₀₀ reading)
- Water bath (for soft agar temperature conditioning)
Before You Start
Determined the phage titer (PFU/mL) of the phage using a plaque assay. A calibration curve relating optical density of the host bacteria at 600 nm (OD₆₀₀) to cell density should be established to enable accurate estimation of bacterial concentration. The host bacterial culture should be grown to the logarithmic phase, corresponding to an OD₆₀₀ of approximately 0.2–0.6, and the CFU/mL at that optical density should be determined. All media required for the experiment should be pre-warmed to the appropriate working temperature to ensure optimal conditions during infection. Sterile dilution tubes should be prepared in advance to facilitate serial dilutions without delays.
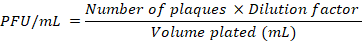
Procedure
One-Step Growth Curve Protocol
Step 1: Preparation of Host Culture
Inoculate the host bacterial strain into LB broth and incubate the culture with continuous shaking until it reaches the mid-logarithmic phase, corresponding to an OD₆₀₀ of approximately 0.2–0.6. Ensure that the culture is actively growing and in an optimal physiological state before proceeding to infection.
Step 2: Infection (Adsorption Phase)
Mix 0.1 mL of phage stock (Assuming titre is ≥ 1×107 PFU/mL) with 9.9 mL host bacterial culture at a multiplicity of infection (MOI) ranging from 0.01 to 0.1. This represents the flask for phage adsorption. (A hundred-fold (1/100) dilution of the original phage suspension yields a final concentration of about 1 × 10⁵ PFU/mL). Incubate the mixture for 5–10 minutes to allow sufficient time for phage adsorption onto the bacterial cells.
Step 3: Removal of Unadsorbed Phages
Centrifuge the infected culture at 10,000 × g for 5 minutes to pellet the bacterial cells along with adsorbed phages. Carefully discard the supernatant containing unadsorbed (free) phages. Resuspend the pellet in fresh 9.9 mL of prewarm medium and repeat the washing step 2–3 times to ensure complete removal of free phages and synchronization of infection.
Step 4: Dilution to Prevent Reinfection
Transfer the defined volume of infected culture into a large volume of prewarmed medium and distribute serially as follows. (Flask A to Flask B to Flask C to Flask D) (Figure 2). Mix thoroughly to achieve a high dilution that prevents secondary infection events from occurring during the experiment. The serial dilution steps should be carried out and completed within a maximum of 2 minutes.
Flask A (Lowest dilution / highest concentration)
Transfer 0.1 mL of infected washed culture into 9.9 mL of LB broth. This represents the initial diluted infected population. (A hundred-fold (1/100) dilution of the original phage suspension yields a final concentration of about 1 × 103 PFU/mL).
Flask B (Intermediate dilution)
Transfer 1.0 mL from Flask A into 9.0 mL fresh medium. (A hundred-fold (1/10) dilution of the original phage suspension yields a final concentration of about 1 × 102 PFU/mL).
Flask C (High dilution)
Transfer 1.0 mL from Flask B into 9.0 mL fresh medium. (A hundred-fold (1/10) dilution of the original phage suspension yields a final concentration of about 1 × 101 PFU/mL).
Flask D (High dilution)
Transfer 1.0 mL from Flask B into 9.0 mL fresh medium. (A hundred-fold (1/10) dilution of the original phage suspension yields a final concentration of about 1 × 10–1 PFU/mL).
This sequential dilution ensures: Reduction of free phage concentration, Prevention of secondary infection, Clear separation of growth kinetics across concentrations.
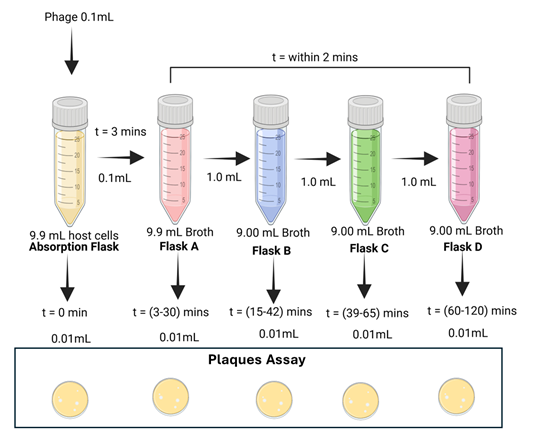
Step 5: Sampling and Plating
Immediately withdraw a 0.1 mL sample from the diluted flask and designate it as time zero (0 minutes). Transfer 0.1 mL from the appropriate flask (A, B, C, or D) according to the overlapping sampling schedule (Table 1), and immediately mix it with 3–4 mL of molten soft agar maintained at approximately 45 °C. Add 200 µL of the host, gently mix, and pour the mixture onto the surface of the underlay agar plate at a slight angle (overlay). Immediately after withdrawing of the 0.1 mL sample from the diluted flasks incubate the flask at 37 °C while shaking for the next time points. Repeat this step promptly and consistently for other time point (3mins, 6mins, 9mins, 12mins, 15 mins …,) to ensure accurate and reliable measurements.
Each plate must be clearly labeled with the corresponding time point and flask (e.g., Flask A, 10 min). After the overlays have solidified (approximately 15–25 minutes), invert the plates and incubate overnight.
If the goal of the first experiment is only to determine the latent period, there is no need to use Flasks C or D. After completing the initial experiment, the sampling frequency and the appropriate flasks to sample from can be better determined.
Step 6: Data Analysis
After incubation, count the number of plaques on each plate. Express all values relative to the phage concentration in Flask A by multiplying counts from the adsorption control (Flask A) by 10, those from Flask B by 100, from Flask C by 1000 and those from Flask D by 10000. Report the final results as PFU/mL.
Plot Log 10 PFU/mL values from the overlapping sampling schedule (Table 1) against time to obtain a growth curve similar to Figure 3, using software such as Microsoft Excel or GraphPad Prism, or by hand on semilogarithmic graph paper.
Table 1: Overlapping Sampling Schedule
| Time (min) | Number of Plaques (Flask A) | Log₁₀ PFU/mL | Number of Plaques (Flask B) | Log₁₀ PFU/mL | Number of Plaques (Flask C) | Log₁₀ PFU/mL | Number of Plaques (Flask D) | Log₁₀ PFU/mL |
| 3 | (30-300) | |||||||
| 6 | (30-300) | |||||||
| 9 | (30-300) | |||||||
| 12 | (30-300) | |||||||
| 15 | (30-300) | (30-300) | ||||||
| 18 | (30-300) | (30-300) | ||||||
| 21 | (30-300) | (30-300) | ||||||
| 24 | (30-300) | (30-300) | ||||||
| 27 | (30-300) | (30-300) | ||||||
| 30 | TMTC (≥300) | (30-300) | ||||||
| 33 | (30-300) | |||||||
| 36 | (30-300) | |||||||
| 39 | (30-300) | (30-300) | ||||||
| 42 | TMTC (≥300) | (30-300) | ||||||
| 45 | (30-300) | |||||||
| 50 | (30-300) | |||||||
| 55 | (30-300) | |||||||
| 60 | (30-300)) | (30-300) | ||||||
| 65 | TMTC (≥300) | (30-300) | ||||||
| 70 | (30-300) | |||||||
| 75 | (30-300) | |||||||
| 80 | (30-300) | |||||||
| 90 | (30-300) | |||||||
| 120 | TMTC (≥300) |
TMTC (≥300) means “Too Many To Count”, indicating that the number of plaques exceeds a reliable counting range (typically ≥300), making accurate enumeration difficult or unreliable.
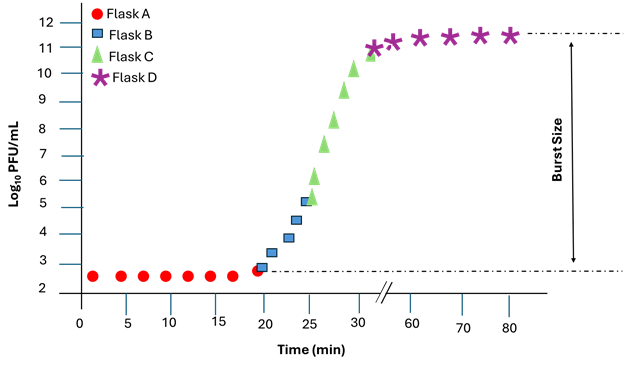
Identify the rise period as the phase characterised by a sharp increase in PFU/mL, indicating phage release. Calculate the burst size using the relation below:
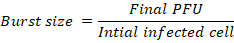
Where:
Final PFU (Plaque Forming Units): This is the total number of infectious phage particles measured at the end of the experiment (usually at the plateau phase of the one-step growth curve).
Initial infected cells: This refers to the number of bacterial cells that were successfully infected by phages at the beginning of the experiment (after adsorption and washing steps). It represents the population of cells that will produce new phages.
Interpretation:
The burst size tells you how many phage particles, on average, are released per infected cell.
For example, if:
- Final PFU = 1,000,000
- Initial infected cells = 100,000
Then:
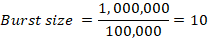
This means each infected bacterial cell produced about 10 phage particles. Burst size is an average value, not an exact count per cell. It reflects the replicative efficiency of the phage under the given experimental conditions.
Results Interpretation
A flat curve is expected during the latent period, followed by a sharp increase corresponding to phage release, and finally a plateau indicating the maximum phage yield. Under properly synchronised conditions, only a single burst should be observed.
Troubleshooting
If multiple bursts are observed, reinfection may have occurred, and the dilution factor should be increased. If low phage recovery is obtained, adsorption may have been inefficient or the phage may be inactive. If the growth curve is unclear, the sampling frequency should be increased. Care should be taken during washing steps to avoid disturbing or losing the bacterial pellet.
Precautions
A low MOI (≤ 0.1) should be maintained to minimise multiple infection events. Washing steps should be performed thoroughly to remove unadsorbed phages. Consistent timing should be maintained across all samples to ensure accuracy. Temperature fluctuations should be avoided throughout the experiment. Strict aseptic techniques should be observed to prevent contamination, and soft agar should not be overheated to avoid damaging phages and host cells.
Applications
This protocol is applied in phage characterisation, infection kinetics studies, evaluation of therapeutic phages, and comparative analysis of phage biology.